
9
September

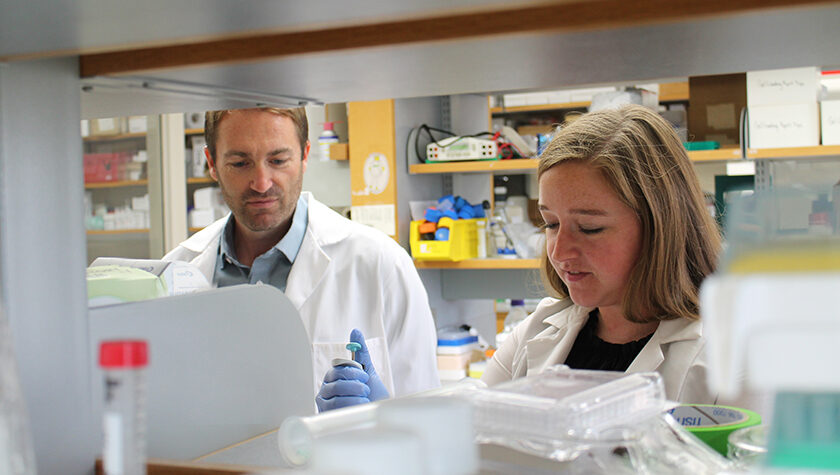
A School of Pharmacy team led by Associate Professor Warren Rose is testing a novel platform that could give doctors a better way to fight hard-to-treat infections
By Nicole Sweeney Etter
A life-threatening infection races through a patient’s bloodstream. But the lab tests to indicate which antibiotic will best defeat the bacteria could take 72 hours — and by then, it could be too late.
A team at the University of Wisconsin–Madison School of Pharmacy is studying a potentially faster, more effective way to test antimicrobial susceptibility, or whether a patient’s bacterial infection will respond to a particular antibiotic. The team includes principal investigator Warren Rose, an associate professor in the Pharmaceutical Sciences and Pharmacy Practice and Translational Research divisions; Sue McCrone, a research specialist in Rose’s lab; and Cecilia Volk (PharmD ’20), an infectious disease fellow and new assistant professor in the Pharmacy Practice and Translational Research Division. Along with colleagues from the UW Carbone Comprehensive Cancer Center, the trio co-authored a paper describing the platform that was published in April in the Royal Society of Chemistry’s Lab on a Chip journal.
“Even if it’s not a serious infection, going back and seeking health care again to get an improved diagnosis or a better antibiotic — those things cost money and time for everyone.”
—Warren Rose
If proven effective, the novel platform could provide hospitals and clinics with a new tool to fight the hardest-to-treat patient infections, Rose says.
“If you’re looking at sepsis or bloodstream infections, the antibiotic failure rate is going be in the 10 to 30 percent range, and a failure would possibly mean death,” he says. “Even if it’s not a serious infection, going back and seeking health care again to get an improved diagnosis or a better antibiotic — those things cost money and time for everyone. And quite frankly, current practice does kind of a poor job of measuring those things.”
Improving antibiotic treatment
Rose’s lab has used modeling systems to simulate human exposure and understand how drugs interact with bacteria. Then Rose heard about under-oil open microfluidic systems (UOMS), a new platform developed by Chao Li, a scientist who works in the lab of David Beebe, a UW professor of pathology and laboratory medicine and biomedical engineering. UOMS cultures cells under oil using very small volumes, more accurately mimicking the environment of the human body.
Li developed UOMS to study interactions with host cells and cancer therapeutics, but Rose wondered if the platform could also be leveraged to test antimicrobial susceptibility.
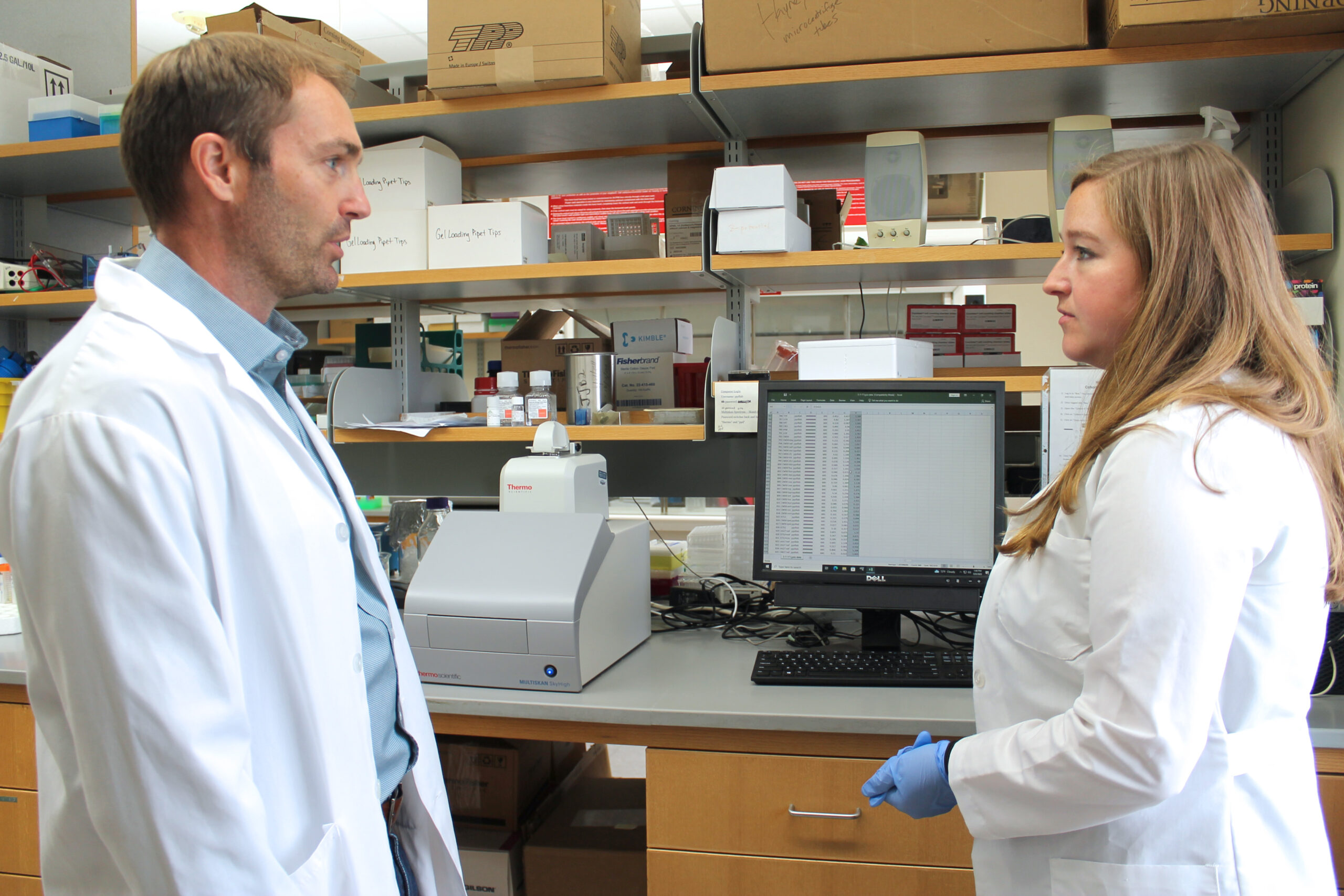
The two labs began collaborating, and the research team discovered that UOMS offers several advantages over culturing methods typically used in hospitals and clinics: Doctors could easily customize antimicrobial susceptibility testing to a patient’s needs, test samples directly from patients instead of pre-culturing samples before putting them into the platform, and test a lot of samples quickly. The platform could also be used to test new antibiotic options in the clinical setting.
“We’re partnering with experts in antimicrobials and engineering technology to test some of these kinds of bad organisms against different types of antibiotics in a very small, niche environment, and we can do it in real time,” Rose says. “The advantage of the platform is to do it not only accurately, which can be done in the clinical lab, but also to do it in very small volumes, with low resources and really low skill needed, and get a turnaround very quickly. You want to know whether this drug is going to work against that pathogen in very acute patients within a few hours.”
The novel platform delivers results in three to four hours — just a fraction of the 24–72 hours usually required.
Researchers believe that the platform can also do something else that traditional methods can’t: detect heteroresistance. That’s when most of a bacterial population succumbs to an antibiotic, but small pockets of resistant bacteria remain, eventually overpowering the drug.
“Studies have shown that it’s more common than we thought,” Rose says. “We’re figuring out that even if resistance is not changing, there’s an increasing underlying population that’s growing more resistant, and the theory is that once a patient starts an antibiotic, then that heteroresistance will emerge as a treatment failure. And if we have the availability to test for this, this could help us understand how and why patients fail.”
The power of collaboration
The project is made stronger by its cross-disciplinary connections.
“It’s really exciting to partner with people you haven’t partnered with before and seeing how they work through things,” says Rose of Li and Beebe.
The colleagues within Rose’s lab also complement each other’s skills. McCrone is the microbiology expert. “She’s been integral,” Rose says. “She understands the culturing techniques and what we’re looking for in organisms.”

McCrone used traditional culturing techniques on Pseudomonas, a common type of bacteria that can cause serious infection, and then ensured that Li used the same conditions to test it on the new platform.
“The results mimicked what was found in the lab in a more efficient way, using less of the sample, antibiotics, media, effort, and time,” she says. “It is also cool to watch a video of the organism multiplying or see no growth when the right antibiotic at the right concentration is given.”
Volk, who previously worked in the Rose Lab as a student and recently returned as a fellow, has studied host-pathogen interactions and is helping the team understand the impact of heteroresistance. She is also helping validate the results using traditional methods and draws on her experience as a clinician to recommend which bacteria and antibiotics they test. She’s especially excited about how quickly the novel platform can deliver results.
“This can lead to faster initiation of appropriate antibiotics for our patients, which has proven to be important in achieving better outcomes,” she explains. “The potential to regularly screen for heteroresistance using a simple, accessible platform will also revolutionize the management of these infections.”
Exploring new possibilities
The initial publication described the novel platform. Now, the research team will continue to test its efficacy and capability.
“We definitely have our eye on this heteroresistance angle and also getting a sense of potentially what interactions might be going on between the host and the pathogen,” Rose explains. “There are ways that we can include host cells in this assay to look at antibiotic activity in context with human cells, which we know impact how antimicrobials and bacteria respond to things. And I think there are ways that we could understand how resistance happens, in terms of novel mechanisms, and ways that we can target niche populations.”
Researchers could also use the platform to co-culture organisms together.
“We have really only begun to discover what this platform can do. It is also ideal for studying host-pathogen interactions and could really push the envelope on personalized medicine in infectious disease management.”
—Cecilia Volk
“Patients sometimes can have mixed infections with different strains, so how those interact and how antibiotics can be tested in that kind of environment would be really interesting,” Rose says.
The research team is excited about the possibilities.
“We have really only begun to discover what this platform can do,” Volk says. “It is also ideal for studying host-pathogen interactions and could really push the envelope on personalized medicine in infectious disease management.”
If further testing confirms that UOMS can outperform existing assays, the platform could be ready for clinical use within five years, Rose says.
“UW has a very long history of antibiotic discovery and development, and I think this just feeds into that history,” Rose says. “And certainly, the resources available here, such as WARF [Wisconsin Alumni Resource Foundation], provide an avenue for reaching patients if the platform is successful.”






